Publications
Preprint
- Ruhela A, Liyanagedra SBW, Laohakunakorn N & Marland JRK (2025) Horseradish peroxidase as an electrochemical reporter protein for cell-free biosensors. bioRxiv https://doi.org/10.64898/2025.12.17.694927
- Stam MJ, Oyarzún DA, Laohakunakorn N & Wood CW (2024) Large scale analysis of predicted protein structures links model features to in vivo behaviour. bioRxiv https://doi.org/10.1101/2024.04.10.588835
Peer-reviewed
- Kulkarni V, Laohakunakorn N & Liyanagedera SBW (2025) Synthetic cells for phage therapy: a perspective. Frontiers in Cellular and Infection Microbiology 15 https://doi.org/10.3389/fcimb.2025.1690404
- Liyanagedera SBW, Bougas A, Yadav S, Wagner C, Zeballos N & Laohakunakorn N (2025) Refinement of OnePot PURE and crude ribosome production for reproducible cell-free protein synthesis. Journal of Visualized Experiments 222:e68826 https://doi.org/10.3791/68826
- Thornton EL, Boyle JT, Laohakunakorn N & Regan L (2025) Cell-free protein synthesis as a method to rapidly screen machine learning-generated protease variants. ACS Synthetic Biology 14:1710-1718 https://doi.org/10.1021/acssynbio.5c00062
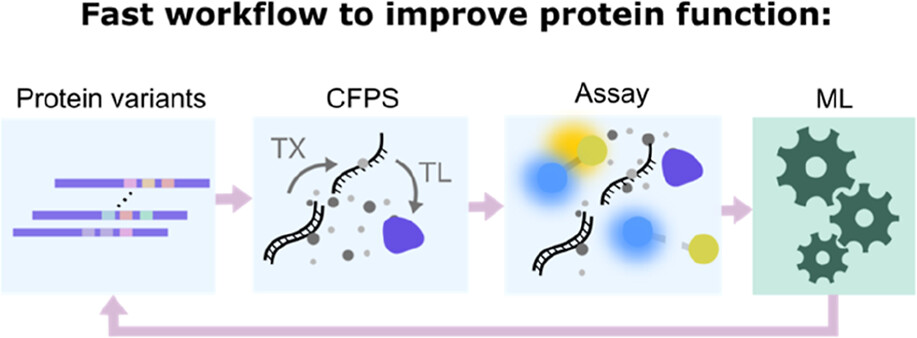
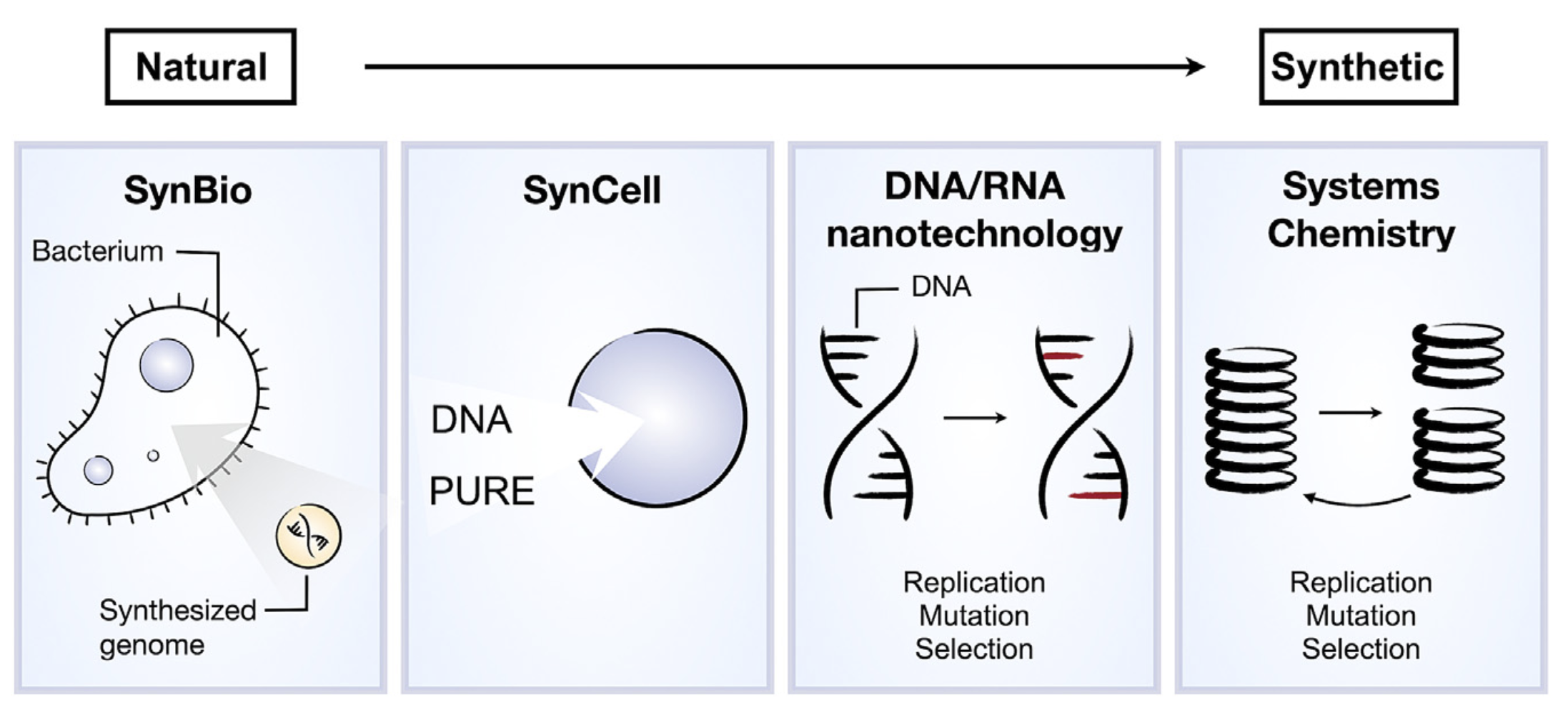
- Kriebisch C et al. (2025) A roadmap towards the synthesis of life Chem 11:102399 https://doi.org/10.1016/j.chempr.2024.102399
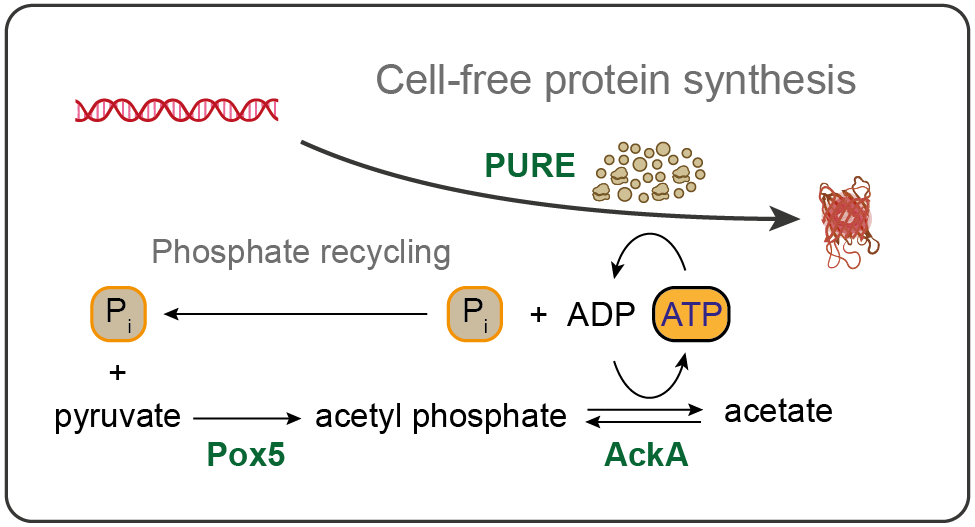
- Yadav S, Perkins AJP, Liyanagedera SBW, Bougas A & Laohakunakorn N (2025) ATP regeneration from pyruvate in the PURE system. ACS Synthetic Biology 14:247-256 https://doi.org/10.1021/acssynbio.4c00697
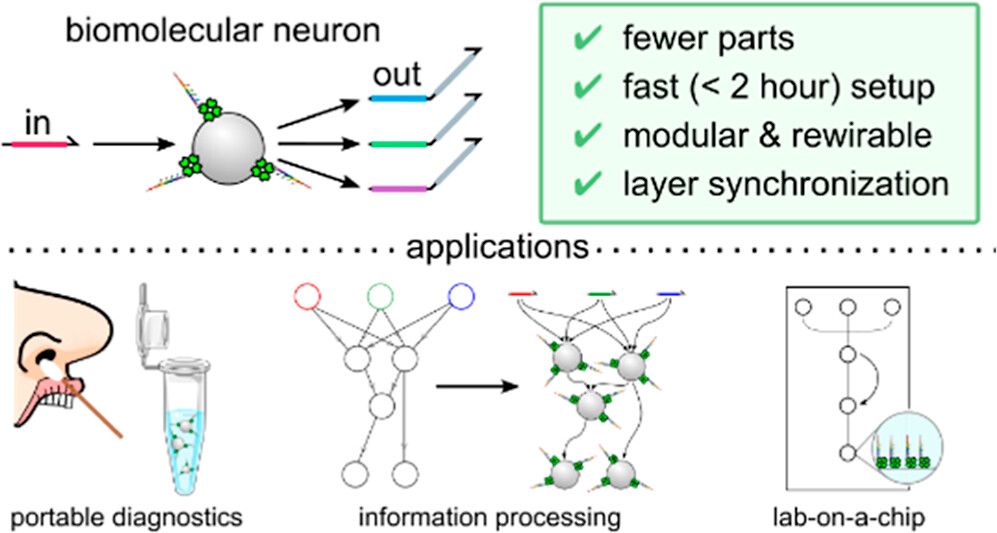
- Lee RC, Corsano A, Tseng CY, Laohakunakorn N & Chou LYT (2024) Rewireable building blocks for enzyme-powered DNA computing networks. Journal of the American Chemical Society 146:26148-26160 https://doi.org/10.1021/jacs.4c07221
- Thornton EL, Paterson SM, Stam MJ, Wood CW, Laohakunakorn N & Regan L (2024) Applications of cell free protein synthesis in protein design. Protein Science 33:9 https://doi.org/10.1002/pro.5148
- Thornton EL, Paterson SM, Gidden Z, Horrocks MH, Laohakunakorn N & Regan L (2022) Self-assembling protein surfaces for in situ capture of cell-free-synthesized proteins. Frontiers in Bioengineering and Biotechnology 10 https://doi.org/10.3389/fbioe.2022.915035
- Laohakunakorn N, Lavickova B, Swank Z, Laurent J & Maerkl SJ (2021) Steady-state cell-free gene expression with microfluidic chemostats. In: Menolascina F. (eds) Synthetic Gene Circuits. Methods in Molecular Biology 2229 https://doi.org/10.1007/978-1-0716-1032-9_9
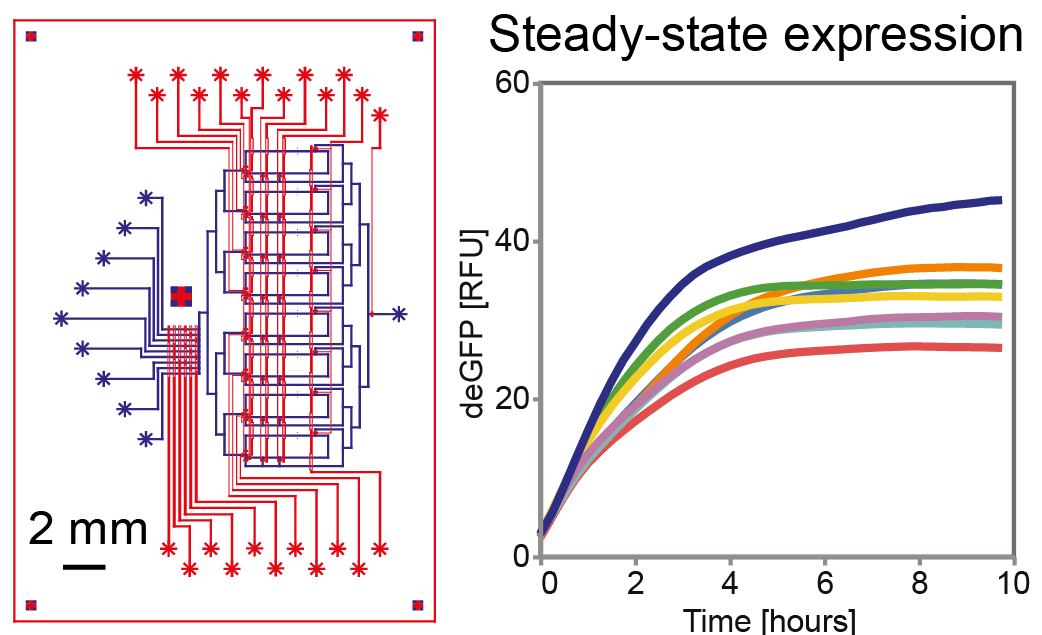
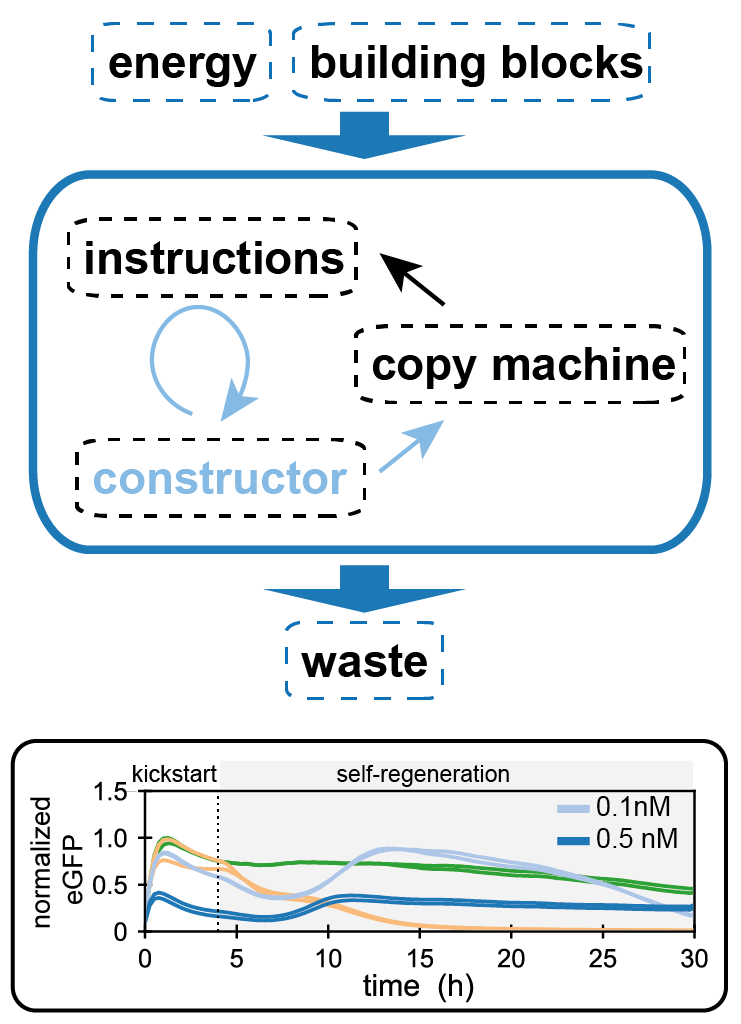
- Lavickova B, Laohakunakorn N & Maerkl SJ (2020) A partially self-regenerating synthetic cell. Nature Communications 11:6340 https://doi.org/10.1038/s41467-020-20180-6 One of the most exciting challenges in future cell-free research is developing systems which can sustainably regenerate their own components. Building such biomolecular 'universal constructors' will elucidate fundamental principles of biochemical autocatalysis, and provide a strategy to power future artificial cells. Here we demonstrate the first steps in using the PURE system to regenerate some of its components, inside controlled microfluidic chemostat reactors. We use our microfluidic platform to assess the regeneration potential of the system, and begin to unravel the critical role that resource allocation and competition play in maintaining robustness of self-regeneration.
- Laohakunakorn N (2020) Cell-free systems: a proving ground for rational biodesign. Frontiers in Bioengineering and Biotechnology 8:788 https://doi.org/10.3389/fbioe.2020.00788
- Laohakunakorn N, Grasemann L, Lavickova B, Michielin G, Shahein A, Swank Z & Maerkl SJ (2020) Bottom-up construction of complex biomolecular systems with cell-free synthetic biology. Frontiers in Bioengineering and Biotechnology 8:213 https://doi.org/10.3389/fbioe.2020.00213
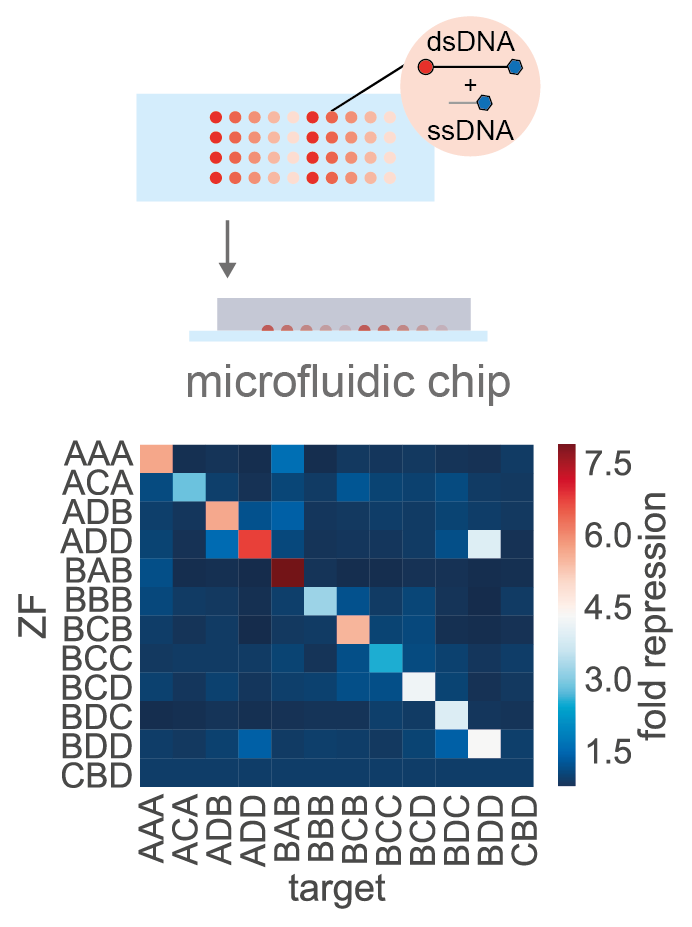
- Swank* Z, Laohakunakorn* N & Maerkl SJ (2019) Cell-free gene regulatory network engineering with synthetic transcription factors. Proc. Natl. Acad. Sci. U.S.A. 116:5892-5901 https://doi.org/10.1073/pnas.1816591116 (* Joint first authors). This paper demonstrates the use of high-throughput, integrated microfluidic devices for characterising the biophysical properties of cell-free gene regulation. We used the device to rapidly characterise a synthetic repressor library, and the results were used to direct the development of strong, cooperative repressors which were assembled to create logic gates in cell-free gene circuits. Featured in EPFL news.
- Rempfer G, Ehrhardt S, Laohakunakorn N, Davies GB, Keyser UF, Holm C & de Graaf J (2016) Selective trapping of DNA using glass microcapillaries. Langmuir 32:8525-8532 https://doi.org/10.1021/acs.langmuir.6b02071
- Laohakunakorn N & Keyser UF (2015) Electroosmotic flow rectification in conical nanopores. Nanotechnology 26:275202 https://doi.org/10.1088/0957-4484/26/27/275202 Conical nanopores exhibit ionic current and electroosmotic flow rectification, both arising from the same phenomenon of ionic permselectivity, which was elucidated in this computational study. Flow rectification is an important function in microfluidic circuits which is typically difficult to achieve due to the linearity of low Reynolds number flows.
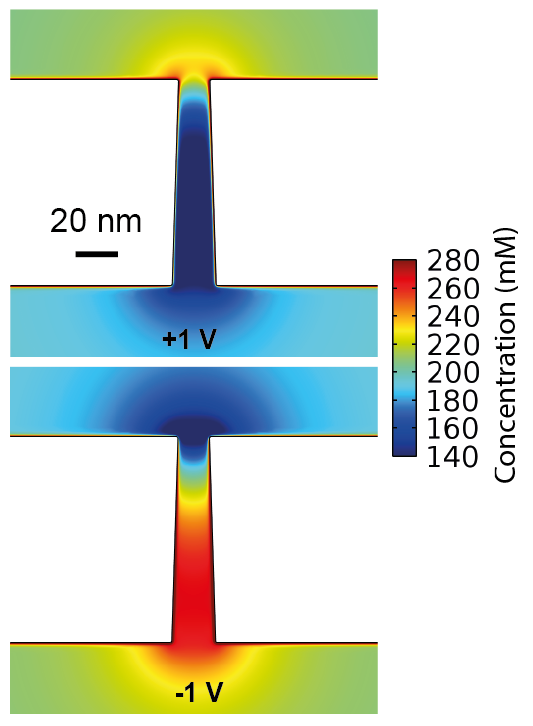
- Laohakunakorn N, Thacker VV, Muthukumar M & and Keyser UF (2015) Electroosmotic flow reversal outside glass nanopores. Nano Letters 15:695-702 https://doi.org/10.1021/nl504237k
- Yanagishima T, Laohakunakorn N, Keyser UF, Eiser E & Tanaka H (2014) Influence of internal viscoelastic modes on the Brownian motion of a lambda-DNA coated colloid. Soft Matter 10:1738-1745 https://doi.org/10.1039/C3SM52830H
- Laohakunakorn N, Otto O, Sturm S, Kroy K & Keyser UF (2013) Dynamic single-molecule force spectroscopy using optical tweezers and nanopores. Proceedings of SPIE 8810:88101F https://doi.org/10.1117/12.2027106
- Laohakunakorn N, Gollnick B, Moreno-Herrero F, Aarts D, Dullens RPA, Ghosal S & Keyser UF (2013) A Landau-Squire nanojet. Nano Letters 13:5141-5146 https://doi.org/10.1021/nl402350a We discovered that an electroosmotic jet driven from the tip of a nanocapillary sets up a large scale fluid flow whose pattern corresponds to a Landau-Squire stokeslet. The mathematical properties of the flow field allowed us to determine precisely the very small flow rate from the pore itself.
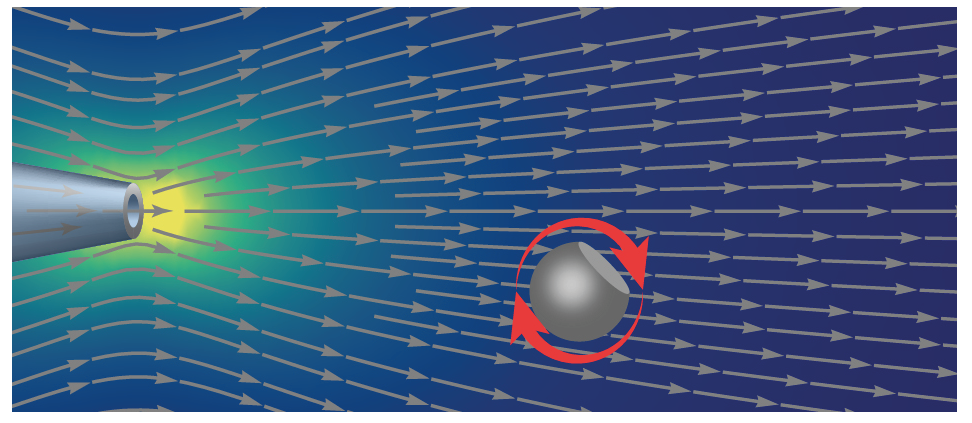

- Laohakunakorn N, Ghosal S, Otto O, Misiunas K & Keyser UF (2013) DNA interactions in crowded nanopores. Nano Letters 13:2798-2802 https://doi.org/10.1021/nl401050m We used single-molecule measurements of DNA molecules tethered between a nanopore and an optical trap to elucidate the electrokinetic forces acting on DNA molecules in confinement, which is important for controlling their translocation through the pore.
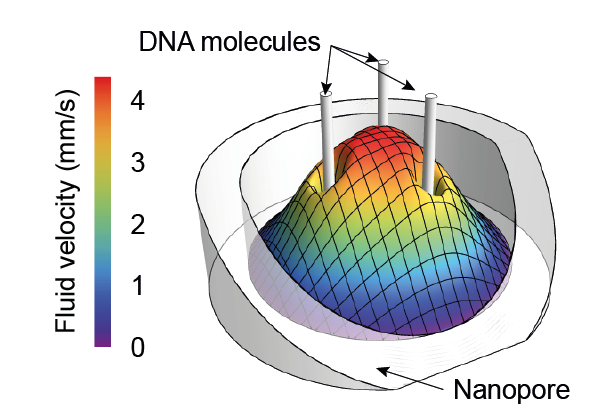
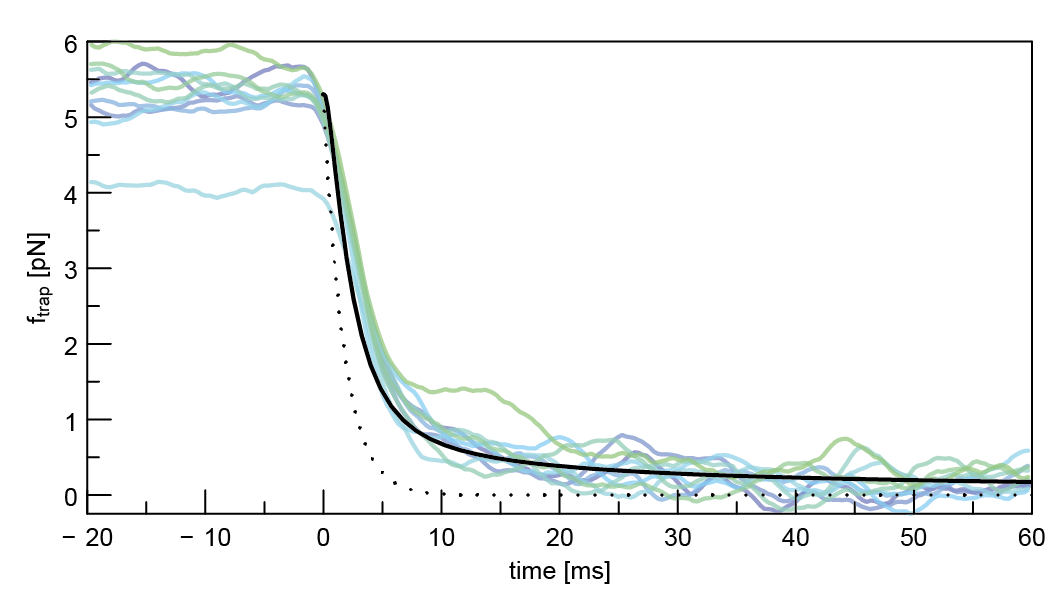
- Otto O, Sturm S, Laohakunakorn N, Keyser UF & Kroy K (2013) Rapid internal contraction boosts DNA friction. Nature Communications 4:1780 https://doi.org/10.1038/ncomms2790 We used the sudden release of DNA tethered between a nanopore and an optical trap to probe its relaxation dynamics, which is dominated by hydrodynamic friction along the length of the molecule. The observations are in agreement with an exact analytical theory.

- Plavchan P, Gueth T, Laohakunakorn N & Parks JR (2013) The identification of 93 day periodic photometric variability for YSO YLW 16A. Astronomy and Astrophysics 554:A110 https://doi.org/10.1051/0004-6361/201220747
Laohakunakorn N (2015) Electrokinetic phenomena in nanopore transport. https://doi.org/10.17863/CAM.16629